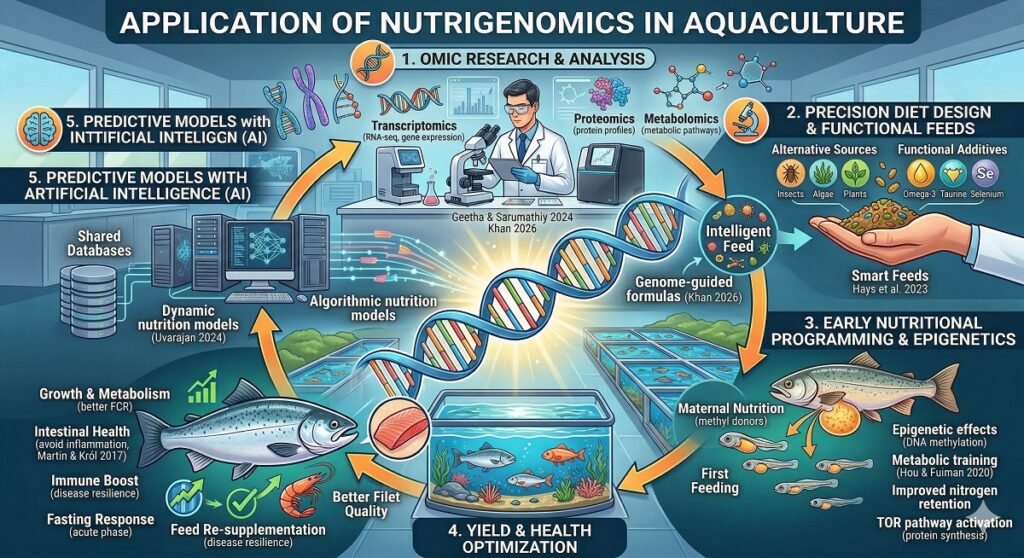
Over the last decade, the aquaculture industry has evolved from an empirical activity into a precision science, with nutrigenomics serving as a fundamental pillar of this transformation. Given the rising global population and the increasing demand for aquatic proteins, developing efficient and sustainable diets has become a strategic imperative. Nutrigenomics seeks not merely to feed a species, but to ‘interact’ with its genes to maximize health, growth, and resilience.
Traditional approaches to feed formulation often overlook the complex interactions between diet, physiology, and the environment (Saluja et al., 2026); in response, nutrigenomics has emerged as a disruptive solution. Today, this discipline has moved beyond the ‘proof of concept’ phase to establish itself as a practical toolkit for designing species-specific functional feeds tailored to biological stages. In this article, we will explore the significance, tools, and challenges of nutrigenomics, integrating key concepts of epigenetics, biotechnology, and functional nutrition.
- 1 Key Takeaways: The Future of Aquaculture Nutrigenomics
- 2 What is Nutrigenomics? Definition and Fundamental Concepts
- 3 Omic Tools: The Arsenal of Modern Aquaculture
- 4 The Molecular Biology of Nutrition: Epigenetics and Gene Expression
- 5 Applications of Nutrigenomics: Transforming Aquaculture Production
- 6 Early Nutritional Programming: The Power of the Beginning
- 7 Immunity, Functional Feeds, and Gut Health
- 8 Sustainability: Plant-Based Proteins and Alternative Oils
- 9 Knowledge Gaps and Future Challenges
- 10 Conclusion: Toward Precision Aquaculture
-
11
Frequently Asked Questions (FAQ) about Aquaculture Nutrigenomics
- 11.1 What is the actual difference between classical nutrition and nutrigenomics?
- 11.2 Can nutrigenomics help reduce production costs?
- 11.3 Is nutrigenomics the same as genetic modification (GMOs)?
- 11.4 What is “nutritional programming” and why is it so important?
- 11.5 How does climate change affect fish nutrition?
- 11.6 Are the benefits of an improved diet passed down to future generations?
- 11.7 Why is Artificial Intelligence discussed in fish nutrition?
- 11.8 What is the difference between nutrigenomics and nutrigenetics?
- 12 References
- 13 Entradas relacionadas:
Key Takeaways: The Future of Aquaculture Nutrigenomics
- From Observation to Precision: The industry has transitioned from ‘trial and error’ methods to a precision science, directly ‘engaging’ with fish and shrimp genomes to optimize performance.
- Nutritional Programming (The ‘Memory Effect’): Nutritional stimuli during larval stages can metabolicly ‘prime’ organisms, enabling carnivorous species to efficiently utilize plant-based diets in adulthood.
- Gut Health and Functional Diets: Transcriptomics allows for the molecular detection of enteritis and metabolic stress before clinical signs appear, facilitating the design of feeds that serve as immunological ‘shields.’
- The Power of Epigenetics: Broodstock nutrition (maternal nutrition) imprints molecular marks on offspring that enhance growth and resilience—a vital advancement for hatchery profitability.
- Convergence with Artificial Intelligence: The near future integrates multi-omic data with AI predictive models to create dynamic feed formulas that adjust in real-time to the animal’s physiological state and environment.
- Economic Efficiency: Precision nutrition transcends science into profitability; reducing culture cycles and improving Feed Conversion Ratios (FCR) transforms the cost structure of any aquaculture operation.
What is Nutrigenomics? Definition and Fundamental Concepts
To grasp the impact of this discipline, it is essential to define the precise meaning of nutrigenomics. In simple terms, it is the study of how nutrients and dietary components influence an organism’s gene expression.
Definition and Technical Scope
Nutrigenomics applies advanced genomic tools—such as transcriptomics—to analyze the nutritional impact on a species’ transcriptome, proteome, and metabolome. In the aquaculture sector, this science investigates how ingredients interact with the genomes of fish and crustaceans to modulate their molecular response. Unlike classical nutrition, this discipline seeks to optimize health, growth, and sustainability through genetic modulation. Several authors support its strategic relevance:
- Hakim et al. (2018): Highlight that nutrigenomics explains why different species process food differently, such as variations in carbohydrate digestion between carnivorous and herbivorous fish.
- Mohanty et al. (2020): Defines its practical objective as understanding individual responses to food to design personalized diets based on genetic makeup.
- Zehra et al. (2025): Notes that it enables the development of species- or strain-specific formulas, satisfying unique metabolic needs through “precision diets.”
- Muhamad et al. (2025): emphasize that this science bridges nutrition and molecular biology to explore how dietary inputs determine phenotypic outcomes in aquatic species.
In practice, this allows us to understand why certain ingredients enhance development, while others may trigger inflammation or metabolic stress processes.
Nutrigenetics vs. Nutrigenomics: Key Differences
While these terms are often confused, they represent two complementary dimensions of molecular nutrition:
- Nutrigenetics (Individual Variability): Focuses on how genetic variations (such as polymorphisms or SNPs) influence the response to diet. For example, it explains why certain Atlantic salmon lineages utilize Omega-3 fatty acids more efficiently than others.
- Nutrigenomics (Response to Stimuli): Studies the inverse effect; that is, how dietary components alter gene expression.
While nutrigenetics helps select individuals with superior metabolic capabilities, nutrigenomics optimizes feed so that these individuals can reach their maximum genetic potential.
Pro-Tip: The synergy between both disciplines allows for a reduction in the culture cycle by aligning the specimen’s genetic profile with a high-precision nutritional program.
Omic Tools: The Arsenal of Modern Aquaculture
The application of nutrigenomic tools is the central axis for addressing critical challenges in larviculture and aquaculture production. According to Gargotra and Aakash (2026), the integration of “omic” technologies provides unprecedented precision regarding nutrient-gene interactions, metabolic pathways, and microbiome dynamics. Furthermore, Saluja et al. (2026) define this approach as an integrative strategy that enables the development of species-specific diets to maximize biological performance.
Stay Always Informed
Join our communities to instantly receive the most important news, reports, and analysis from the aquaculture industry.
The “Omics” Discipline Ecosystem
Based on research by Chuphal and Singha (2026), Iyyapparaja et al. (2022), and Mohanty et al. (2020), the key contributions of each discipline are presented below:
| Discipline | Function and Application in Aquaculture |
| Genomics | Identifies genetic variations impacting metabolism. Uses DNA sequencing to map the complete fish genome. |
| Transcriptomics | Demonstrates how gene expression varies with different diets. RNA-seq provides insights into larval growth and immunity. |
| Proteomics | Analyzes the protein set (proteome). Identifies growth biomarkers and key proteins associated with production. |
| Metabolomics | Studies cellular chemical reactions. Considered the final product of transcription, it reveals health and feed efficiency. |
| Epigenomics | Explains how diet induces heritable changes without altering DNA, enabling “nutritional programming” in early stages. |
Genome Editing Technologies: The Potential of CRISPR/Cas9
Genome editing via CRISPR/Cas9 is redefining aquaculture biotechnology. While nutrigenomics adjusts the “software” (gene expression), CRISPR allows for editing the “hardware” (the genetic code) to enhance desirable traits. According to Khan (2026), integrating CRISPR with nutrition will enable disruptive applications:
- Carnivorous fish with herbivorous enzymes: Enabling carnivorous species to digest plant-based ingredients, eliminating dependence on fishmeal.
- Shielded microbiomes: Designing pathogen-resistant gut bacteria to protect fish naturally.
- Next-Generation Editing: Tools like Prime Editing and Base Editing offer surgical precision to correct individual letters of the genetic code without damaging the DNA strand.
Legal Disclaimer: The use of genetically edited organisms is subject to strict international regulations. Consumer acceptance currently remains the greatest challenge for commercial implementation.
Key Advancements and Future Applications
Technological convergence is accelerating industry results (according to Ibrahim et al., 2025 and Zhang et al., 2025):
- Input Substitution: Development of strains capable of endogenous synthesis of Omega-3 fatty acids (EPA and DHA), reducing fish oil usage.
- Reference Genomes: Access to high-quality genomes (salmon, trout, tilapia) facilitates the discovery of specific nutrigenomic markers.
- Predictive Bioinformatics: Advanced workflows allow for models that predict the relationship between diet, immunity, and health before the culture cycle begins.
The Molecular Biology of Nutrition: Epigenetics and Gene Expression
Nutrigenomics in fish and crustaceans relies heavily on epigenetics—the study of heritable changes in gene function that do not alter the original DNA sequence, but rather its execution.
DNA Methylation and Growth Regulation
One of the most extensively studied mechanisms in teleost fish is DNA methylation. Biochemically, the addition of a methyl group to cytosine residues within dinucleotides is typically associated with the silencing of specific genes. This process can be represented by the following enzymatic interaction:
In modern aquaculture, nutritional factors such as methionine or choline levels can alter these methylation patterns. This directly impacts genes responsible for growth regulation and resilience to environmental stress.
The Role of mRNA and Transcriptomics
Transcriptomics stands as the master tool of nutrigenomics. By quantifying messenger RNA () levels, researchers determine which genes are activated or deactivated in response to a diet. For instance, if a species receives excessive levels of plant protein, transcriptomics can reveal the up-regulation of pro-inflammatory genes in the gut, alerting to the risk of enteritis before physical symptoms become visible.
Current technologies have enabled significant milestones, according to experts:
- Geetha and Sarumathiy (2024): Report that Next-Generation Sequencing (), combined with bioinformatic tools such as DESeq2 and EdgeR, allows for the discovery of genes that are differentially expressed in response to minimal nutritional changes.
- Khan (2026): Classifies DNA microarrays and as essential “omic” technologies for analyzing transcription outputs and understanding nutrient influence at the molecular level.
Genomic Diagnostic Tools
According to scientific evidence, these tools are applied in the aquaculture field as follows:
- DNA Microarrays: Offer a comprehensive view of the genomic response by analyzing thousands of genes simultaneously. Although powerful, Khan (2026) notes that their use still faces limitations due to the need for species-specific genetic markers.
- RNA-seq: This advanced technology allows for the observation of how components like Omega-3 fatty acids can attenuate cytokine excess during viral infections by up-regulating interferon genes and Natural Killer () cells to combat pathogens.
Applications of Nutrigenomics: Transforming Aquaculture Production
The transition from a “trial and error” model toward precision nutrition represents the most disruptive advancement of the decade. According to Khan (2026), current applications allow for production optimization across four strategic axes:
Metabolic Optimization Strategies
- Species Differentiation: Nutrigenomics explains why carnivorous fish face difficulties assimilating carbohydrates due to insulin dysregulation. In contrast, it identifies how herbivorous fish activate specific enzymes to process these inputs, facilitating more cost-effective diets.
- Protein and Lipid Efficiency: By reducing protein and increasing lipids through “genomically guided” methods, protein synthesis is enhanced while nitrogen waste is minimized. This protects fillet quality and reduces environmental pollution without causing excessive visceral fat accumulation.
- Immunological Precision Diets: Specific nutrients act as genetic “switches.” For instance, n-3 PUFAs (Omega-3) reduce viral load and inflammation, while selenium hyper-stimulates natural antiviral defenses.
- Designer Feeds: The ultimate goal is to maximize the Feed Conversion Ratio (FCR) and accelerate compensatory growth, thereby shortening the culture cycle and reducing environmental impact.
Overcoming Hatchery Challenges and Environmental Stress
Early mortality and cannibalism are often symptoms of nutritional imbalances invisible to the human eye.
- Iyyapparaja et al. (2022) emphasize that omic tools are vital for understanding how nutrition affects cellular response during critical developmental stages.
- Hays et al. (2023) demonstrated that precise supplementation (such as 1.2% taurine in turbot) positively stimulates growth genes and enriches essential metabolic pathways.
- Mitra et al. (2023) highlight a vital application in the face of the climate crisis: dietary modification to mitigate physiological stress from temperature changes, strengthening natural immunity through gene expression.
Summary of Advancements by Biological Traits
| Trait / Objective | Nutrigenomic Advancement | Key References |
| Input Substitution | Transcriptomics guides the use of plant proteins (soy, wheat, pea) without affecting muscle metabolism. | (Dhanasiri et al., 2020; Ibrahim et al., 2025) |
| Health & Immunity | Omics reveal how functional feeds modulate critical pathways such as NF-κB and JAK-STAT. | (Martin & Król, 2017; Kari, 2025) |
| Protein Sparing | Optimization of the protein-lipid-carbohydrate ratio for more efficient energy utilization. | (Exadactylos, 2014; Saha et al., 2020) |
| Crustacean Performance | Linking nutrients with molting, growth, and antioxidant defense genes in shrimp. | (Alhoshy et al., 2022; Hossain et al., 2020) |
| Nutritional Programming | Early dietary management positively alters the intestinal transcriptome for the fish’s entire lifespan. | (Rimoldi et al., 2025; Martin & Król, 2017) |
Early Nutritional Programming: The Power of the Beginning
A disruptive concept in the industry is Early Nutritional Programming. This phenomenon demonstrates that stimuli received during larval stages can “program” a fish’s metabolism for the rest of its life, optimizing its performance through adulthood.
Training Metabolism from Birth
Metabolic programming consists of using specific larval diets to “precondition” enzymatic pathways. According to Hou and Fuiman (2020), this developmental plasticity coincides with periods of high mortality, allowing nutrition to be employed as a strategic tool for survival and sustainability.
Kumar et al. (2026) summarize the competitive advantages of this approach:
- Permanent Effects: Interventions during embryonic, larval, and juvenile phases induce physiological and neurological changes that persist for a lifetime.
- Maternal Inheritance: Nutrients transferred from the mother via the yolk sac dictate initial viability. Substituting fish oils with vegetable oils in the maternal diet can “teach” offspring to better tolerate and process these alternative sources in the future.
- The Power of First Feeding: The first external feed shapes digestive capacity. A brief early exposure to carbohydrates leads to a prolonged increase in the activity of enzymes such as amylase, enabling carnivorous species to efficiently assimilate plant-based diets in adulthood.
The Epigenetic Engine and the TOR Pathway
The mechanism behind this metabolic memory is epigenetics. Alterations such as DNA methylation and histone modification adapt the fish’s phenotype to its environment without altering its genetic code.
In larviculture, the clearest example is the use of specific amino acids to activate the TOR (Target of Rapamycin) signaling pathway, which is responsible for protein synthesis. Hakim et al. (2018) emphasize that these effects are so profound that they can even be transmitted to subsequent generations, impacting overall growth and immune response.
Advancements in Larval Development
Despite knowledge gaps regarding neurological development, technologies such as are enabling the design of specific micro-diets.
- Iyyapparaja et al. (2022): Note that genetic precision will allow for the creation of tailored diets to maximize hatchery survival.
- Martínez-Burguete et al. (2021): Identified for the first time the transcriptomic profiles of the tropical gar (Atractosteus tropicus), establishing candidate markers to monitor digestive system development.
Impact of Nutritional Components on Development
| Nutritional Component | Observed Genetic Effect | Productive Benefit |
| Retinoic Acid | Regulation of bone development genes. | Drastic reduction in malformations. |
| Methionine | Global DNA methylation. | Significant improvement in nitrogen retention. |
| Prebiotics | Modulation of intestinal mucin genes. | Increased natural resistance to pathogens. |
Immunity, Functional Feeds, and Gut Health
Immune function in aquatic organisms is intrinsically linked to their nutrition. Nutrigenomics has uncovered how functional feeds modulate biological responses at a molecular level, transforming disease prevention into a precision science.
Modulation of the Immune Response
According to Martin and Król (2017), the strategic use of additives—such as prebiotics (beta-glucans and oligosaccharides), phytochemicals, and specific marine oils—enhances natural resistance. Khan (2026) reinforces this concept, reporting that nutrigenomics enables the exact identification of how these components regulate gene expression within both innate and adaptive immune pathways.
The Challenge of Plant-Based Proteins and Oils
The transition toward plant-derived ingredients (soy, pea) due to fishmeal scarcity has presented critical challenges that transcriptomics helps mitigate:
- Intestinal Inflammation: Certain anti-nutritional factors in plants trigger adverse responses. Transcriptomic studies cited by Martin and Król (2017) reveal that soy rapidly activates inflammatory mediator genes, which can lead to intestinal epithelial barrier dysfunction, thereby facilitating pathogen entry.
- Fasting Management: Nutrient deprivation reduces the expression of key immune system components to conserve energy. However, fasting fish exhibit a disproportionate “acute-phase” response to infections in an attempt to compensate for diminished prior defenses. Furthermore, fasting significantly weakens the mucosal immune system in the skin and gills.
Strategic Comparison: Traditional Nutrition vs. Nutrigenomic Approach
| Feature | Traditional Nutrition | Nutrigenomic Approach |
| Primary Objective | Survival and final weight. | Molecular health and gene efficiency. |
| Tools | Growth trials (FCR). | Transcriptomics, Proteomics, and Metabolomics. |
| Precision | Low (final phenotype observation). | High (prior cellular response observation). |
| Ingredients | Based on availability and cost. | Optimized to avoid genomic inflammation. |
| Sustainability | Dependent on marine inputs. | High (facilitates alternative protein use). |
Sustainability: Plant-Based Proteins and Alternative Oils
The transition toward fishmeal-free diets represents the greatest challenge in modern aquaculture. In this scenario, nutrigenomics is positioned as the definitive tool for understanding why carnivorous species develop pathologies such as enteritis when ingesting plant proteins, enabling the selection of ingredients that do not trigger inflammatory pathways.
Challenges of the Nutritional Transition
The use of vegetable oils and plant proteins is an ecological and economic necessity. However, these inputs contain anti-nutritional factors (ANFs) that can compromise the integrity of the intestinal epithelium. Nutrigenomics allows for the mapping of these adverse responses to formulate diets that mitigate cellular damage. According to the most recent research:
- Gargotra and Aakash (2026): Strategies based on “omic” disciplines facilitate the adoption of sustainable alternatives, such as insect meal and algal biomass, fostering the efficient use of non-marine resources.
- Zamzami et al. (2025): By decoding metabolic pathways, it is possible to maintain production performance in shrimp using plant or microbial protein sources. This reduces dependence on finite inputs and supports the development of low-impact aquaculture systems aligned with global SDGs (Sustainable Development Goals).
Toward an “Herbivorous Metabolism” in Carnivores
One of the most disruptive advancements is the use of nutritional programming to “teach” species such as salmon to process carbohydrates efficiently. Although their genome is not naturally designed for this purpose, manipulating gene expression from early stages allows for a forced metabolic adaptation, optimizing the use of terrestrial inputs in traditionally carnivorous species.
Knowledge Gaps and Future Challenges
Despite the disruptive potential of fish nutrigenomics, the industry faces critical hurdles that must be overcome to achieve widespread and standardized implementation:
- Causality vs. Correlation: It is fundamental to move beyond observing which genes are activated to understand the biological “why,” determining if such changes are the direct drivers of observed growth.
- Lack of Full-Cycle Studies: Most current research is short-term. Longitudinal studies are required to monitor organisms from the larval stage through to harvest.
- Standardization of Protocols: Variability in results between laboratories highlights the need for unified methods for epigenetic assays in aquatic species.
- Antinutrient Synergy: A knowledge gap persists regarding how multiple anti-nutritional factors (ANFs) interact at the molecular level when combined in a single diet.
The Future: Artificial Intelligence and Precision Nutrition
The evolution of the discipline points toward massive data integration. According to Uvarajan (2024), combining genomic, proteomic, and metabolomic levels will feed Artificial Intelligence (AI) predictive models. This will allow for the formulation of diets that dynamically adapt to the physiological state of each species, maximizing profitability and minimizing nutrient waste. Strategic directions for 2025–2026 include:
- Precision Nutrition Frameworks: Integration of genotype and environment to design “tailored feeds” (Ibrahim et al., 2025; Zhang et al., 2025).
- Nutritional Immunomodulation: Utilizing nutrigenomics to optimize immunostimulants, seeking total resilience and the elimination of antibiotics (Kari, 2025).
- Multi-omic Integration: Identifying durable nutritional markers through the study of non-coding RNA (ncRNA).
Pending Challenges and Global Vision
As noted by Zehra et al. (2025) and Gargotra & Aakash (2026), the future lies in personalized feeding strategies that ensure global food security. However, to get there, we must address three pillars:
- Multigenerational Studies: Confirming whether the epigenetic benefits of broodstock diets are stably inherited.
- Fillet Quality: Linking gene expression to the final texture, color, and nutritional profile of the commercial product.
- Regulatory and Social Framework: There is an urgent need to educate policymakers and consumers on the safety and benefits of these biotechnologies.
Conclusion: Toward Precision Aquaculture
Nutrigenomics in aquaculture has evolved from an academic trend into the master key for sustainable “blue production.” By integrating nutritional science with molecular genetics, it is now possible to develop diets that not only maximize growth but also guarantee animal welfare and drastically reduce the industry’s environmental footprint.
This discipline is laying the groundwork for a new generation of “smart feeds” driven by omic data. As noted by Zamzami et al. (2025), these formulas synergistically integrate antioxidant regulation, immunological conditioning, and precision engineering of the gut microbiome.
Ultimately, nutrigenomics has displaced the outdated “trial and error” model in favor of an approach based on exact biological mechanisms. Although challenges remain regarding long-term effects and species-specificity, the sector’s future lies in our ability to decipher the dialogue between nutrients and the genome. As we bridge current technical gaps, aquaculture will undoubtedly position itself as the most efficient, sustainable, and technologically advanced animal protein production sector on the planet.
Frequently Asked Questions (FAQ) about Aquaculture Nutrigenomics
What is the actual difference between classical nutrition and nutrigenomics?
While classical nutrition focuses on observing external outcomes—such as weight gain or survival rates—nutrigenomics analyzes cellular and genetic responses. This allows for the detection of underlying issues, such as subclinical intestinal inflammation, long before the fish exhibits stunted growth or mortality.
Can nutrigenomics help reduce production costs?
Yes. By utilizing omic tools, formulators can substitute expensive ingredients (such as fishmeal) with more cost-effective alternatives (plant-based or insect meals) without compromising animal health. Furthermore, early nutritional programming can shorten the culture cycle, leading to direct savings in energy and labor costs.
Is nutrigenomics the same as genetic modification (GMOs)?
No. This is a common misconception. Nutrigenomics studies how feed influences the genes an organism already possesses (adjusting the ‘software’). In contrast, genetic modification or CRISPR editing alters the DNA code itself (changing the ‘hardware’). While both disciplines can work in tandem, nutrigenomics alone does not create Genetically Modified Organisms (GMOs).
What is “nutritional programming” and why is it so important?
It is a technique where larvae are exposed to specific nutrients to ‘train’ their metabolism. For example, a carnivorous salmon can be programmed from birth so that, as an adult, it digests plant proteins as efficiently as an herbivorous fish. This is the key to low-impact aquaculture.
How does climate change affect fish nutrition?
Thermal stress caused by warming waters alters gene expression and weakens the immune system. Nutrigenomics enables the design of ‘resilience diets’ with functional additives that protect cells from oxidative damage and maintain productivity even under adverse environmental conditions.
Are the benefits of an improved diet passed down to future generations?
Yes, through epigenetics. It has been demonstrated that broodstock nutrition (maternal nutrition) leaves ‘marks’ on the DNA that do not change the sequence itself, but rather activate growth and health genes in the larvae—providing them with a competitive advantage even before birth.
Why is Artificial Intelligence discussed in fish nutrition?
Due to the vast amount of data generated by transcriptomics and proteomics, AI and Machine Learning models are now utilized to process this information. This enables the creation of ‘feed recipes’ that change dynamically based on water temperature, the fish’s age, and its specific genetic profile.
What is the difference between nutrigenomics and nutrigenetics?
Nutrigenomics studies the effect of diet on genes, whereas nutrigenetics examines how a fish’s genes determine its response to that diet.
References
Alhoshy, M., Shehata, A., Habib, Y., Abdel‐Latif, H., Wang, Y., & Zhang, Z. (2022). Nutrigenomics in crustaceans: Current status and future prospects. Fish & shellfish immunology. https://doi.org/10.1016/j.fsi.2022.08.056
Chuphal, N., Singha, K.P. (2026). Nutrigenomics for Sustainable Aquaculture. In: Ishfaq Nazir, M., Ahmad, I. (eds) Aquatic Genomics and Food Security . Springer, Cham. https://doi.org/10.1007/978-3-032-14971-8_2
Dhanasiri, A., Johny, A., Xue, X., Berge, G., Bogevik, A., Rise, M., Fæste, C., & Fernandes, J. (2020). Plant-Based Diets Induce Transcriptomic Changes in Muscle of Zebrafish and Atlantic Salmon. Frontiers in Genetics, 11. https://doi.org/10.3389/fgene.2020.575237
Exadactylos, A. (2014). Nutrigenomics in Aquaculture Research. Fisheries and Aquaculture Journal, 5. https://doi.org/10.4172/2150-3508.1000e107
Gargotra, P., Aakash, M. (2026). Future Outlook: The Impact of Omics on the Future of Aquaculture Nutrition. In: Ishfaq Nazir, M., Ahmad, I. (eds) Aquatic Genomics and Food Security . Springer, Cham. https://doi.org/10.1007/978-3-032-14971-8_17
Geetha, K., & Sarumathiy, C. K. (2024). Frontiers in Marine Biotechnology and Fish Genomics: Innovations for Sustainable Aquaculture and Blue Economy Advancement. National Journal of Smart Fisheries and Aquaculture Innovation, 2(2), 58-63. https://doi.org/10.17051/NJSFAI/02.02.08
Hakim, M. M., Ganai, N. A., Ahmad, S. M., Asmi, O., Akram, T., Hussain, S., Shabir, N., Shah, R., & Gora, A. H. (2018). Nutrigenomics: Omics approach in aquaculture research to mitigate the deficits in conventional nutritional practices. Journal of Entomology and Zoology Studies, 6(4), 582-587
Hays, H., Gu, Z., Mai, K., & Zhang, W. (2023). Transcriptome-based nutrigenomics analysis reveals the roles of dietary taurine in the muscle growth of juvenile turbot (Scophthalmus maximus). Comparative Biochemistry and Physiology Part D: Genomics and Proteomics, 47, 101120. https://doi.org/10.1016/j.cbd.2023.101120
Hossain, M., Koshio, S., & Kestemont, P. (2020). Recent advances of nucleotide nutrition research in aquaculture: a review. Reviews in Aquaculture. https://doi.org/10.1111/raq.12370
Hou, Z., Fuiman, L.A. Nutritional programming in fishes: insights from mammalian studies. Rev Fish Biol Fisheries 30, 67–92 (2020). https://doi.org/10.1007/s11160-019-09590-y
Houston, R., Bean, T., Macqueen, D., Gundappa, M., Jin, Y., Jenkins, T., Selly, S., Martin, S., Stevens, J., Santos, E., Davie, A., & Robledo, D. (2020). Harnessing genomics to fast-track genetic improvement in aquaculture. Nature Reviews Genetics, 21, 389 – 409. https://doi.org/10.1038/s41576-020-0227-y
Ibrahim, S., Yue, C., Yang, C., Ran, Y., Xie, B., Yang, Q., Mkuye, R., Liang, H., & Deng, Y. (2025). A review of genomics application in aquaculture: status, challenges, and priorities for future direction. Annals of Animal Science, 0. https://doi.org/10.2478/aoas-2025-0078
Kari, Z. (2025). Nutritional immunomodulation in aquaculture: functional nutrients, stress resilience, and sustainable health strategies. Aquaculture International, 33. https://doi.org/10.1007/s10499-025-02122-5
Khan, A. (2026). Nutrigenomics in Aquaculture: Decoding Gene–Diet Interactions for Sustainable Fish Production. Agrolife Frontiers, 1(1), 9-16.
Kumar, Shivendra, Banik, Aditi, Dubey, Maneesh Kumar, Srivastava, Prem Prakash, Sandor, Zsuzsanna J., Early Nutritional Programing: Unlocking the Potential of Fish for Sustainable Aquaculture, Aquaculture Nutrition, 2026, 3380126, 25 pages, 2026. https://doi.org/10.1155/anu/3380126
Iyyapparaja Narasimapallavan, G., Sanal Ebeneezar, Vijayagopal, P., Linga Prabu, D., Sajina, K. A., Krupesha Sharma, S. R., Sumithra, T. G., Tamilmani, G., Saktivel, M., Anikuttan, K. K., & Gopalakrishnan, A. (2022). Nutrigenomics tools to address the emerging issues in marine fish larviculture. En D. Dayal & M. Shamsudheen (Eds.), Advances in Agricultural, Animal and Fisheries Sciences (pp. 20-29). ZNAN Publishers, Society for Technology, Environment, Science & People. https://doi.org/10.5281/zenodo.6473509.
Martin, S. A., & Król, E. (2017). Nutrigenomics and immune function in fish: New insights from omics technologies. Developmental & Comparative Immunology, 75, 86-98. https://doi.org/10.1016/j.dci.2017.02.024
Martínez-Burguete, T., Peña-Marin, E. S., García-Gasca, A., Alvarez-González, C. A., & Llera-Herrera, R. (2021). Nutrigenomic marker discovery by de novo transcriptomic sequencing during early development of the tropical gar (Atractosteus tropicus). Aquaculture Research, 52(8), 3829-3842. https://doi.org/10.1111/are.15228
Mitra, A., Abdel-Gawad, F. K., Bassem, S., Barua, P., Assisi, L., Parisi, C., Temraz, T. A., Vangone, R., Kajbaf, K., Kumar, V., & Guerriero, G. (2023). Climate Change and Reproductive Biocomplexity in Fishes: Innovative Management Approaches towards Sustainability of Fisheries and Aquaculture. Water, 15(4), 725. https://doi.org/10.3390/w15040725
Mohanty, B. P., Ganguly, S., Mahanty, A., Mitra, T., & Mohanty, S. (2020). Nutrigenomics and fish. CAB Reviews, 15(048). https://doi.org/10.1079/PAVSNNR202015048
Muhamad Firdaus, Muhammad Dailami, Desy Setyoningrum, Muhammad Arif Rahman. Fish Nutrigenomics and Molecular Strategies for Sustainable Aquaculture. Fish Nutrigenomics and Molecular Strategies for Sustainable Aquaculture, 1-103, BP International, 2025, ?10.9734/bpi/mono/978-93-49970-87-8?. ?hal-05144568?
Munang’andu, H. M., & Evensen, Ø. (2017). Current Advances in Functional Genomics in Aquaculture. In Applications of RNA-Seq and Omics Strategies – From Microorganisms to Human Health. InTech. https://doi.org/10.5772/intechopen.69883
Rimoldi, S., Quiroz, K., Kalemi, V., McMillan, S., Stubhaug, I., Martinez-Rubio, L., Betancor, M., & Terova, G. (2025). Interactions between nutritional programming, genotype, and gut microbiota in Atlantic salmon: Long-term effects on gut microbiota, fish growth and feed efficiency. Aquaculture. https://doi.org/10.1016/j.aquaculture.2024.741813
Saha, A., Nayak, M., & Giri, S. (2020). Nutrigenomics Studies in Fishes. **, 215-228. https://doi.org/10.1201/9781003107583-7
Saluja, S. et al. (2026). Nutritional Security Through Advanced Omics Technologies in Aquaculture. In: Ishfaq Nazir, M., Ahmad, I. (eds) Aquatic Genomics and Food Security . Springer, Cham. https://doi.org/10.1007/978-3-032-14971-8_8
Uvarajan KP, Omics-Driven Insights into Nutrigenomics and Growth Optimization in Aquatic Species, National Journal of Smart Fisheries and Aquaculture Innovation, Vol. 2, No. 1, 2024 (pp.62-68).
Zamzami, I. M. A., Prihanto, A., Mohamad Lal, M. T., & Jaziri, A. (2025). Global Research Trends in Genome-Based Aquafeed for Pacific White Shrimp (Litopenaeus vannamei): A Scientometrics Analysis (1976–2025). Egyptian Journal of Aquatic Biology and Fisheries, 29(5), 3305-3327. doi: 10.21608/ejabf.2025.434435.6842
Zehra, S., Alghamdi, M., Mohamed, A.H., Alhafedh, Y.S., Al-Shaikhi, A. (2025). Role of Genomics and Nutrigenomics in Aquaculture Nutrition. In: Ahmed, I., Ahmad, I. (eds) Aquaculture: Enhancing Food Security and Nutrition. Springer, Cham. https://doi.org/10.1007/978-3-031-92858-1_11
Zhang, Y., Lu, R., Qin, C., & Nie, G. (2020). Precision nutritional regulation and aquaculture. Aquaculture Reports, 18, 100496. https://doi.org/10.1016/j.aqrep.2020.100496
Zhang, J., Li, M., Meng, D., Xu, S., Teame, T., Yao, Y., Yang, Y., Zhang, Z., Ran, C., Jijakli, M., Ding, Q., & Zhou, Z. (2025). Review on Omics Approaches in Aquatic Animal Nutrition: Current Status, Limitations, and Perspectives. The Journal of nutrition. https://doi.org/10.1016/j.tjnut.2025.08.019
Editor at the digital magazine AquaHoy. He holds a degree in Aquaculture Biology from the National University of Santa (UNS) and a Master’s degree in Science and Innovation Management from the Polytechnic University of Valencia, with postgraduate diplomas in Business Innovation and Innovation Management. He possesses extensive experience in the aquaculture and fisheries sector, having led the Fisheries Innovation Unit of the National Program for Innovation in Fisheries and Aquaculture (PNIPA). He has served as a senior consultant in technology watch, an innovation project formulator and advisor, and a lecturer at UNS. He is a member of the Peruvian College of Biologists and was recognized by the World Aquaculture Society (WAS) in 2016 for his contribution to aquaculture.