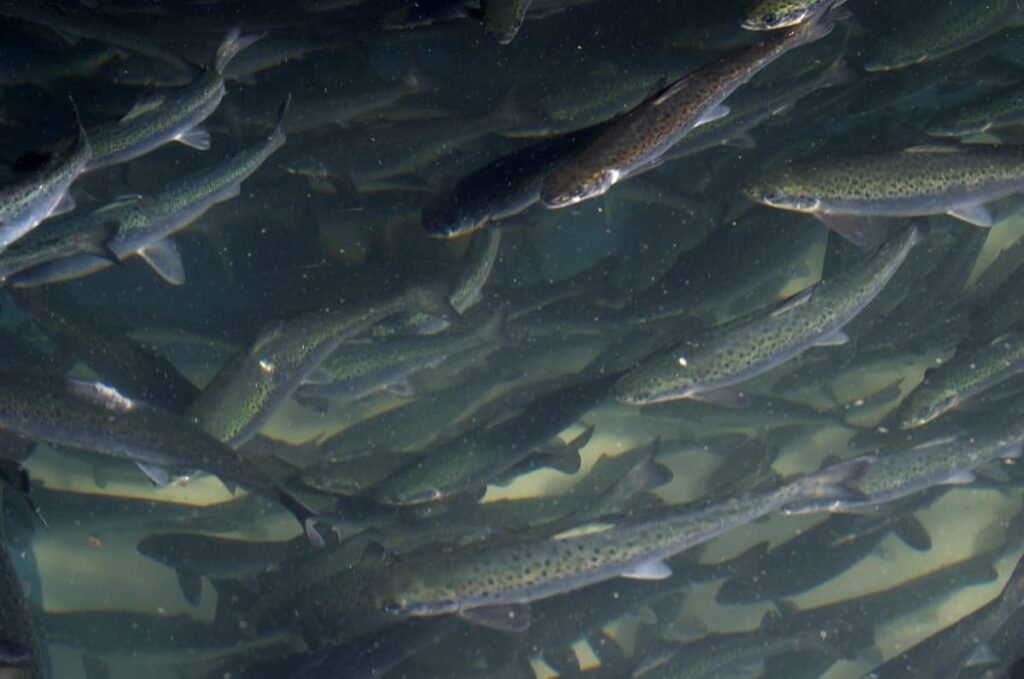
Atlantic salmon (Salmo salar L.) is one of the most valuable commodities in global trade, with production reaching 2.86 million tonnes in 2022, led by Norway. However, this industry faces a persistent and silent enemy: Listeria monocytogenes. This pathogen is responsible for listeriosis, an infection with an alarming mortality rate of 19.7% in the European Union.
Despite massive efforts to sanitize processing plants, outbreaks linked to smoked salmon and ready-to-eat (RTE) products continue to occur. Until now, science had focused on factory cleanliness, but a new study published in the journal Aquaculture (2026) proposes a disruptive perspective: What if the problem originates from the feed?
Key Findings
- Hidden Transmission Route: A direct genetic link was identified between Listeria strains found in feed and those in salmon processing plants.
- Diversity at the Source: The highest variety of sequence types (ST) of the bacteria was concentrated in feed mills, rather than in the sea or processing facilities.
- Pathogen Persistence: Strains ST8 and ST37, known for their ability to survive in industrial environments, were the only ones detected in the final processing stage.
- Paradigm Shift: The results suggest that Listeria control must begin long before the fish reaches the slaughterhouse, focusing instead on the feed supply chain.
Tracing the Genetic Footprint in 1,800 Samples
To solve this enigma, a team of scientists from NTNU (Norwegian University of Science and Technology) and the University of Copenhagen designed a risk-based sampling plan covering the entire production chain for nearly a year.
The Study Map:
- Feed Mills: Three facilities (FF-A, FF-B, FF-C) and the vessels transporting the feed.
- Marine Phase: Three open-sea salmon farms, including samples from healthy fish and natural mortalities.
- Transport and Processing: Well-boats and primary processing plants where salmon is converted into fillets.
In total, 1,819 samples were analyzed using cutting-edge techniques such as Oxford Nanopore-based repetitive extragenic palindromic PCR (ON-rep-seq) and Whole Genome Sequencing (WGS) to differentiate strains with surgical precision.
Feed Mills as Reservoirs of Diversity
The results revealed an overall L. monocytogenes prevalence of 2.7%. While this figure seems low, the data distribution tells a revealing story:
- Feed Chain: 8% positivity.
- Marine Phase: Only 1%.
- Processing Plant: 3%.
The most striking finding was not the quantity, but the genetic diversity. Ten different sequence types (ST) were identified; of these, eight were present in the feed production chain. In contrast, only two or three STs were detected in the marine phase and the processing plant.
The “Smoking Gun”: ST8 and ST37
The study found that strains ST8 and ST37 exhibited extreme genetic similarity (fewer than 10 allelic differences in the core genome) between feed mill samples and the final product at the processing plant. This connection strongly suggests that the pathogen can travel through the feed or its logistical environment to reach the final consumer.
The Extrusion Process: An Insufficient Shield
Interestingly, feed extrusion (where temperatures exceed 100°C) appears to eliminate Listeria effectively, as no bacteria were detected immediately after this step. However, recontamination occurs rapidly during storage in outdoor bulk bags, ship transport, or distribution through piping at marine farms. Factors such as humidity in silos and exposure to birds or pests at loading docks were identified as critical risk points.
Implications for Industry and Public Health
This study marks a turning point in aquaculture biosecurity. Currently, regulations do not mandate feed producers to perform exhaustive Listeria testing, focusing almost exclusively on Salmonella.
Stay Always Informed
Join our communities to instantly receive the most important news, reports, and analysis from the aquaculture industry.
The Importance of Persistence
The detected strains, especially ST8, are known “persistent clones” capable of surviving for years in hostile environments by forming biofilms and developing resistance to disinfectants. If these strains are continuously introduced through feed, cleaning efforts in filleting plants will be, at best, a temporary solution.
Limitations and Next Steps
Despite the solid genetic evidence, researchers note that detection at sea is difficult due to water dilution. Furthermore, while it is suspected that the bacteria may colonize the salmon’s gut via contaminated feed, intestinal content analysis was not performed in this phase—a key recommendation for future research. Science must now determine how frequently Listeria from feed survives the fish’s digestive tract and whether this constitutes the primary entry point into slaughterhouse evisceration machinery.
Contact
Sunniva Hoel
NTNU – Norwegian University of Science and Technology, Department of Biotechnology and Food Science
7491 Trondheim, Norway
Email: sunniva.hoel@ntnu.no
Reference (open access)
Thomassen, G. M. B., Jakobsen, A. N., Lerfall, J., Jameson, J., Krych, L., Hannisdal, A., Knøchel, S., & Hoel, S. (2026). Tracing Listeria monocytogenes in the Atlantic salmon (Salmo salar L.) production chain – from feed to primary processing. Aquaculture, 616, 743733. https://doi.org/10.1016/j.aquaculture.2026.743733
Editor at the digital magazine AquaHoy. He holds a degree in Aquaculture Biology from the National University of Santa (UNS) and a Master’s degree in Science and Innovation Management from the Polytechnic University of Valencia, with postgraduate diplomas in Business Innovation and Innovation Management. He possesses extensive experience in the aquaculture and fisheries sector, having led the Fisheries Innovation Unit of the National Program for Innovation in Fisheries and Aquaculture (PNIPA). He has served as a senior consultant in technology watch, an innovation project formulator and advisor, and a lecturer at UNS. He is a member of the Peruvian College of Biologists and was recognized by the World Aquaculture Society (WAS) in 2016 for his contribution to aquaculture.