Biodiversity monitoring in aquatic ecosystems has undergone a profound metamorphosis thanks to environmental DNA (eDNA), a technique that identifies species by analyzing the genetic traces they leave behind. However, the precision of this biological “fingerprint” has been hindered by incomplete databases, contaminated sequences, and species that appear genetically identical.
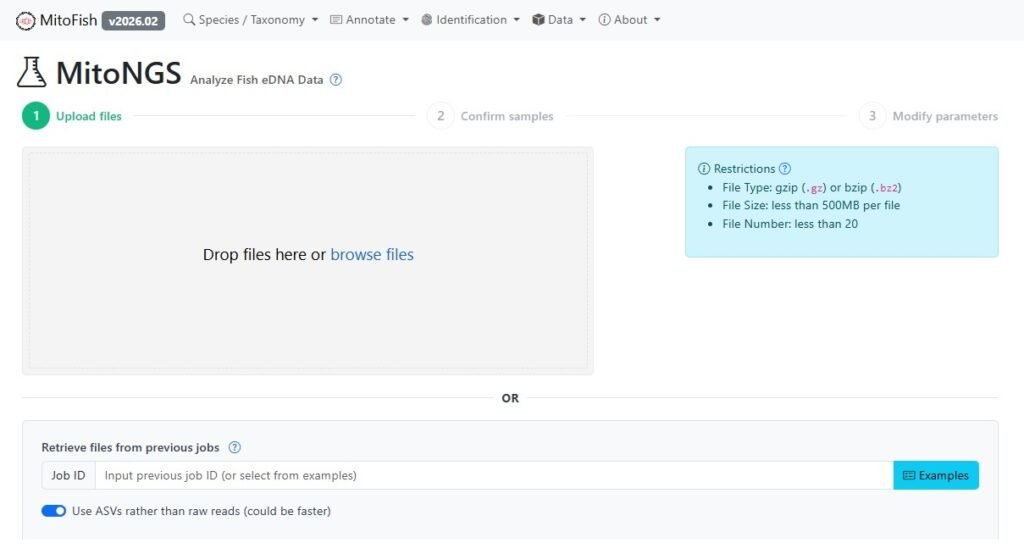
To address these bottlenecks, a research team led by Tao Zhu and Susumu Yoshizawa from The University of Tokyo has launched MitoNGS, a next-generation web platform succeeding the highly successful MiFish, which introduces unprecedented contextual intelligence into marine bioinformatics.
- 1 Key Study Conclusions
- 2 Key Differences: Why MitoNGS Surpasses MiFish
- 3 Strategies for High-Precision Taxonomic Identification
- 4 Global Impact and Results: From Japan to the Amazon
- 5 Challenges and Future Perspectives in Genomic Biomonitoring
- 6 Strategic Relevance in Aquatic Resource Management
- 7 Conclusion: The New Standard in Global Biomonitoring
- 8 Entradas relacionadas:
Key Study Conclusions
- Full Compatibility: MitoNGS analyzes data from any mitochondrial marker () and sequencing platforms such as Illumina and Nanopore.
- Resolution via “Species Complexes”: It introduces a strategy to manage taxa with identical sequences by utilizing habitat and geographic distribution data to filter results.
- Contaminated Data Cleansing: The platform identifies and excludes “heterospecific” (mislabeled) regions from reference databases to prevent false positives.
- Real-Time Efficiency: It is significantly faster than competing tools, processing complex Nanopore libraries within minutes.
- Accessible Interface: It offers a free, user-friendly web solution for researchers and biological resource managers.
Key Differences: Why MitoNGS Surpasses MiFish
The superiority of MitoNGS lies in its advanced processing engine and the robustness of its reference database, MitoFish. While the original MiFish pipeline is limited to universal rRNA primers, MitoNGS is a comprehensive platform that supports diverse mitochondrial loci. This versatility is critical for the biotechnology industry, as it allows for the use of markers such as or according to specific research requirements.
Validation of Heterospecific Regions
A critical challenge within GenBank is the proliferation of chimeric sequences: DNA fragments where the target species and contaminants coexist. To mitigate this, MitoNGS fragments mitochondrial genes into 100 bp segments and validates them against homologs from various authors. If a region matches only distinct species, it is classified as heterospecific and excluded from the analysis to ensure the integrity of the results.
The “Species Complex” Approach
Conventional systems often fail when two species present an identical metabarcoding marker ( identity). MitoNGS resolves this by grouping these taxa into a species complex. For precise resolution, the system cross-references information with external variables:
- Habitat Parameters: Salinity, depth, and climate zone (via rFishBase).
- Geographic Distribution: Global occurrence records (via GBIF).
Optimization for Third-Generation Sequencing (Nanopore)
Unlike its predecessor, MitoNGS is optimized for the long and complex reads of platforms like Nanopore. It utilizes the VSEARCH algorithm to generate high-fidelity Amplicon Sequence Variants (ASVs) from raw data, facilitating real-time environmental monitoring through portable devices such as the MinION.
Strategies for High-Precision Taxonomic Identification
One of the most critical challenges in metabarcoding is sequential ambiguity, a phenomenon that occurs when a DNA sequence presents a match with multiple species. To mitigate this obstacle, MitoNGS implements advanced discrimination protocols:
Taxonomic Validation of Subregions
The system decomposes sequences into 100 bp (base pair) fragments to rigorously identify cross-contamination or pre-existing mislabeling errors within GenBank. This process enables the detection of heterospecific regions, ensuring that taxonomic assignment is not compromised by poorly curated sequences in public databases.
Integration of Environmental and Geographic Metadata
The platform transcends pure bioinformatics by integrating ecological variables from rFishBase (depth, salinity, and climate zone) and global occurrence records from GBIF.
Stay Always Informed
Join our communities to instantly receive the most important news, reports, and analysis from the aquaculture industry.
In scenarios where two species share an identical genomic identity, the system discerns the correct taxonomy through biogeographic data cross-referencing. For instance, if one species inhabits Arctic ecosystems while the other resides in tropical regions, the software prioritizes assignment based on the geographic coordinates of the collected sample.
Global Impact and Results: From Japan to the Amazon
The efficacy of the MitoNGS platform has been validated through eight heterogeneous datasets, spanning from the Uchidomari River in Japan to the Peruvian Amazon and various aquarium systems in the Netherlands. This geographic diversity underscores the system’s versatility across distinct aquatic ecosystems.
Performance and Analytical Precision
In testing phases, MitoNGS demonstrated significant technical superiority over competing solutions. During the analysis of a Nanopore library with 2 kb reads, the system completed processing in just 8 minutes, contrasting sharply with the more than 2 hours required by the DECONA pipeline. Furthermore, the platform successfully identified 15 additional species that had gone undetected in the original studies. This increase in taxonomic sensitivity is due to the robust coverage of its database, which is subject to constant updates.
Resolution of Genetic Conflicts
MitoNGS has shown a superior capacity to resolve taxonomic ambiguities. The system allowed for the assignment of specific identities to specimens originally reported as “general clades.”
A notable case is the precise identification of Caracanthus unipinna on Europa Island. This level of resolution is possible because the MitoFish database is updated monthly, ensuring that researchers always work with the most recent genomic information available.
Challenges and Future Perspectives in Genomic Biomonitoring
Despite its technological sophistication, MitoNGS acknowledges and addresses the challenges intrinsic to the current state of environmental genomics. These limitations define the roadmap for future innovations:
- Reference Sequence Gaps: Critical voids persist in uncommon fish families, such as Scytalinidae or Lacantuniidae, for which mitochondrial sequences are not yet available in global databases.
- Hybridization Complexity: The presence of natural hybrids in certain species complexes can blur genetic boundaries. This phenomenon poses a challenge for taxonomic resolution in specific markers where differentiation is minimal.
- Bacterial DNA Interference: When using markers like , the system may detect sequences of bacterial origin. To mitigate this background signal, a continuous expansion of reference databases is required to filter non-target material more accurately.
Strategic Relevance in Aquatic Resource Management
MitoNGS transcends the definition of a simple software tool to establish itself as an integrated data ecosystem. By allowing the upload of custom ASVs/OTUs for dynamic re-analysis, the platform drives a more collaborative, transparent, and reproducible science. For industry professionals, the tool optimizes the management of multiple samples by natively integrating:
- Controls and Replicates: Visual management of positive and negative controls, as well as biological replicates.
- Error Mitigation: Proactive identification of laboratory cross-contamination, drastically reduces the risk of false positives.
- Auditable Results: Data presentation in analytical tables that facilitate critical decision-making.
This technical robustness ensures that reports on invasive species or the health status of fishery stocks are fully reliable and meet the highest standards of scientific auditing.
Conclusion: The New Standard in Global Biomonitoring
MitoNGS establishes itself as the gold standard for fish metabarcoding analysis, achieving an unprecedented synergy between speed, precision, and a web interface designed for the democratization of genomics. By successfully resolving the challenge of ambiguous sequences and optimizing compatibility with cutting-edge technologies like Nanopore, the platform guarantees a more robust and reliable monitoring of global aquatic biodiversity.
This ecosystem breaks traditional industry barriers: today, any researcher with a laptop and an internet connection can execute highly complex analyses. What once required supercomputers and command-line bioinformatics experts is now accessible to the global scientific community, facilitating more agile, data-driven environmental management.
This research was supported by: Ocean DNA Project (No. 027) from Future Society Initiative, Interdisciplinary Collaborative Research Program (1-B) of Atmosphere and Ocean Research Institute, the University of Tokyo, and MEXT Advancement of Technologies for Utilizing Big Data of Marine Life JPMXD 1521474594.
Contact
Tao Zhu
Department of Integrated Biosciences, Graduate School of Frontier Sciences, The University of Tokyo.
Department of Natural Environmental Studies, Graduate School of Frontier Sciences, The University of Tokyo.
Correo: zhutao@edu.k.u-tokyo.ac.jp
Susumu Yoshizawa
Department of Natural Environmental Studies, Graduate School of Frontier Sciences, The University of Tokyo.
Atmosphere and Ocean Research Institute, The University of Tokyo.
Correo: yoshizawa@aori.u-tokyo.ac.jp
Reference (open access)
Zhu, T., Sato, Y., Fukunaga, T., Miya, M., Iwasaki, W., & Yoshizawa, S. (2026). MitoNGS: an online platform to analyze fish metabarcoding data in high resolution. Molecular Biology and Evolution, 43(3), 1-15. https://doi.org/10.1093/molbev/msag046
Editor at the digital magazine AquaHoy. He holds a degree in Aquaculture Biology from the National University of Santa (UNS) and a Master’s degree in Science and Innovation Management from the Polytechnic University of Valencia, with postgraduate diplomas in Business Innovation and Innovation Management. He possesses extensive experience in the aquaculture and fisheries sector, having led the Fisheries Innovation Unit of the National Program for Innovation in Fisheries and Aquaculture (PNIPA). He has served as a senior consultant in technology watch, an innovation project formulator and advisor, and a lecturer at UNS. He is a member of the Peruvian College of Biologists and was recognized by the World Aquaculture Society (WAS) in 2016 for his contribution to aquaculture.