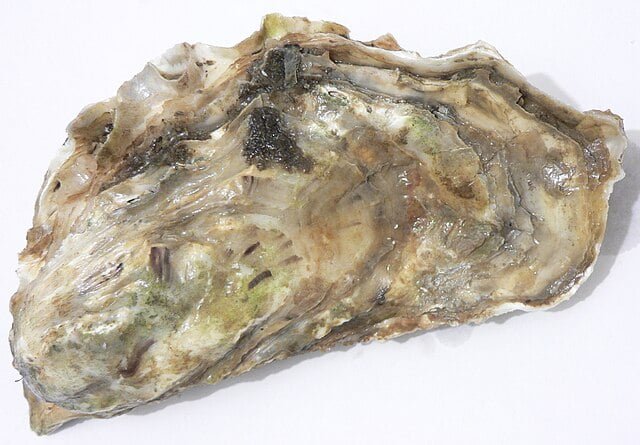
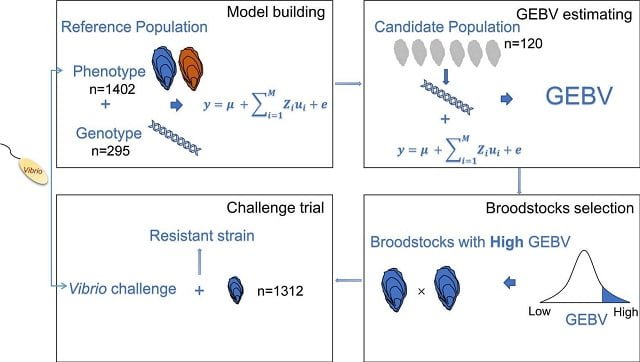
Pacific oyster cultivation relies mainly on collecting seeds from the wild or producing them in hatcheries. Considering its economic potential, selective breeding programs for oysters are being encouraged internationally.
Mass selection has been carried out since the mid-1990s in different oyster species, but it has been widely used to mitigate mortality caused by ostreid herpesvirus 1 (OsHV-1) in Crassostrea gigas.
Although effective, mass selection can quickly lead to inbreeding if genetic diversity is not adequately controlled.
The emergence of genomic tools has allowed new approaches to the use of genomic selection in the design of oyster breeding programs, which are easy and inexpensive to implement compared to classical selective breeding schemes.
A team of scientists from SYSAAF, Ifremer, France Naissain, SATMAR, Université Montpellier, INRAE, evaluated the potential of genomic selection for growth, meat content, and color traits in two selected breeding families of Pacific oyster (Crassostrea gigas).
The researchers first characterized the genetic structure and diversity of the two populations. They then estimated genetic parameters for commercial traits.
Genomic Selection
According to the researchers, “Genomic selection is particularly relevant for traits that are costly or difficult to measure (e.g., disease resistance, meat quantity) because fewer phenotypic data are needed to obtain similar accuracies of resulting EBVs compared to pedigree- and GS-based selection.”
Recent advances in genotyping tools in mollusks have so far resulted in a relatively limited number of studies on the potential of genomic selection.
Genomic selection has been investigated in the Portuguese oyster for morphometric traits, edibility traits, and disease resistance traits, in the American oyster (Crassostrea virginica) for low salinity tolerance, and in the silver-lip pearl oyster for pearl quality traits (Pinctada maxima), among others.
Stay Always Informed
Join our communities to instantly receive the most important news, reports, and analysis from the aquaculture industry.
High Marker Heterogeneity
The study aimed to further demonstrate the feasibility of genomic selection on a commercial scale in the design of mixed families, using two independent populations under selection in France.
According to the study results, there is a strong marker density heterogeneity between and within chromosomes, with a low linkage disequilibrium ratio between pairs of SNPs.
“Growth traits were generally highly genetically and positively correlated with each other but weakly correlated with color traits,” they reported.
They also highlighted that prediction accuracy was generally higher with the genomic model (GBLUP) than with the classical BLUP model, with a maximum gain in accuracy (from 0.38 to 0.66) for meat weight adjusted for total weight at P2.
Conclusion
“Our results confirm the possibility of improving growth and color characteristics in commercial populations of Pacific oysters and the relevance of genomic selection and mixed-family breeding designs to achieve this,” the scientists concluded.
They also emphasized that the two evaluated breeding programs showed substantial genetic variation for genetic improvement and good genetic diversity, as indicated by the estimates of effective size.
However, the researchers recommend developing better genomic tools because the number of high-quality SNPs was low and yielded poor results.
The study was funded by the European Maritime and Fisheries Fund (EMFF) and FranceAgriMer as part of the “Quality Huitre.”
Reference
Antoine Jourdan, Romain Morvezen, Florian Enez, Pierrick Haffray, Adeline Lange, Emilie Vétois, François Allal, Florence Phocas, Jérôme Bugeon, Lionel Dégremont, Pierre Boudry. 2023. Potential of genomic selection for growth, meat content and colour traits in mixed-family breeding designs for the Pacific oyster Crassostrea gigas, Aquaculture, Volume 576, 2023, 739878, ISSN 0044-8486, https://doi.org/10.1016/j.aquaculture.2023.739878.
Editor at the digital magazine AquaHoy. He holds a degree in Aquaculture Biology from the National University of Santa (UNS) and a Master’s degree in Science and Innovation Management from the Polytechnic University of Valencia, with postgraduate diplomas in Business Innovation and Innovation Management. He possesses extensive experience in the aquaculture and fisheries sector, having led the Fisheries Innovation Unit of the National Program for Innovation in Fisheries and Aquaculture (PNIPA). He has served as a senior consultant in technology watch, an innovation project formulator and advisor, and a lecturer at UNS. He is a member of the Peruvian College of Biologists and was recognized by the World Aquaculture Society (WAS) in 2016 for his contribution to aquaculture.