Selective breeding has been successfully implemented in a large number of aquaculture species, and the recent development of genotyping tools, such as high-density single nucleotide polymorphism (SNP) polymorphism, has opened the door to the implementation of genomic selection for most species.
Selective breeding using genomic selection has the potential to achieve cumulative improvements for host resistance. However, genomic selection is expensive due to the cost of genotyping a large number of animals using high-density single nucleotide polymorphism (SNP) arrays.
In this regard, a team of researchers from The Roslin Institute and Royal at the University of Edinburgh, the Natural Resources Institute Finland (Luke), and Benchmark Genetics evaluated the efficiency of genomic selection for Flavobacterium columnare resistance using low-density (LD) in silico panels combined with imputation.
Columnaris disease
Flavobacterium columnare is the pathogenic agent of columnaris disease, an emerging disease that affects rainbow trout (Oncorhynchus mykiss) aquaculture.
The bacterium F. columnare is distributed worldwide and affects freshwater fish, usually when water temperatures are between 18 to 20°C, but it has also been reported to affect salmonids in cold water conditions.
F. columnare causes acute and chronic infections, with main symptoms being tissue and gill necrosis, especially in small fish, leading to high mortality if the disease is not treated.
Low-density SNP Panels (LD)
Several strategies are being investigated to reduce genotyping costs, making genomic selection a more accessible tool for small and medium-scale selective breeding programs.
Many studies have explored the potential of using LD SNP panels for genomic selection in various aquaculture species and have concluded that LD panels contain between 1000 and 6000 SNPs.
According to the study authors, depending on the species and trait, LD panels are sufficient to achieve genomic prediction accuracy similar to that obtained with a medium or high-density panel (HD).
Stay Always Informed
Join our communities to instantly receive the most important news, reports, and analysis from the aquaculture industry.
Main study results
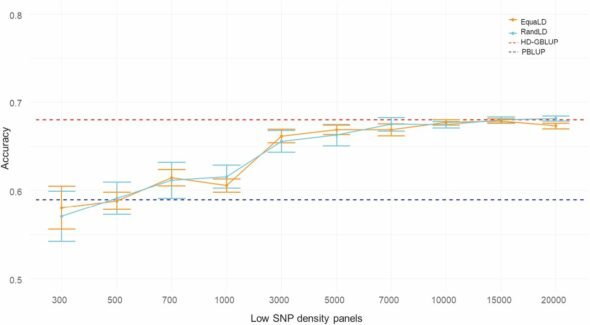
The scientists evaluated the impact of decreasing SNP density on genomic prediction using LD SNP panels created in silico using three sampling methods.
As the researchers report, “The accuracy of imputation and genomic predictions was similar with random and equidistant SNP sampling. The use of LD panels of at least 3000 SNPs or panels of lower density (as low as 300 SNPs) combined with imputation resulted in accuracies comparable to the 28K HD panel and 11% higher than pedigree-based predictions.”
They highlight that their results confirm that for rainbow trout, the accuracy of genomic prediction can be achieved with a low-density marker in the range of 3000 to 7000.
Cost efficiency of genotyping strategies
According to the researchers, “the use of a low-density panel to genotype rainbow trout and genomically predict EC resistance would result in only a small reduction in prediction accuracy (3%) compared to using a high-density panel, while significantly reducing costs (around 25%).”
However, they caution that the hypothetical price of €15 per sample for genotyping using a 3K SNP panel is unrealistic, as such panels do not exist for rainbow trout and could actually be more expensive than estimated, thus reducing the appeal of low-density genotyping.
The study analyzed the relevance of using low-density panels for genomic selection as a way to reduce genotyping costs with a marginal loss of accuracy, allowing small and medium-scale aquaculture breeding programs to implement genomic selection.
Conclusions
“Compared to the use of commercial HD panels, LD panels combined with imputation can provide a more affordable approach to genomic prediction of genetic values, supporting a broader adoption of genomic selection in aquaculture breeding programs,” the researchers concluded.
Contact
Clémence Fraslin
The Roslin Institute and Royal (Dick)
School of Veterinary Studies, University of Edinburgh,
Easter Bush, Midlothian, EH25 9RG, UK
Email: clemence.fraslin@roslin.ed.ac.uk
Reference (open access)
Fraslin, C., Robledo, D., Kause, A. et al. Potential of low-density genotype imputation for cost-efficient genomic selection for resistance to Flavobacterium columnare in rainbow trout (Oncorhynchus mykiss). Genet Sel Evol 55, 59 (2023). https://doi.org/10.1186/s12711-023-00832-z
Editor at the digital magazine AquaHoy. He holds a degree in Aquaculture Biology from the National University of Santa (UNS) and a Master’s degree in Science and Innovation Management from the Polytechnic University of Valencia, with postgraduate diplomas in Business Innovation and Innovation Management. He possesses extensive experience in the aquaculture and fisheries sector, having led the Fisheries Innovation Unit of the National Program for Innovation in Fisheries and Aquaculture (PNIPA). He has served as a senior consultant in technology watch, an innovation project formulator and advisor, and a lecturer at UNS. He is a member of the Peruvian College of Biologists and was recognized by the World Aquaculture Society (WAS) in 2016 for his contribution to aquaculture.